Reproducibility Evaluation (MITRE Study) Reproducibility assessment of an automated multiplexed immunofluorescence slide staining, imaging, and analysis workflow
Authors: Clifford Hoyt 1, Kristin Roman 1, Liz Engle 2, Chichung Wang 1, Carmen Ballesteros-Merino 3, Shawn M. Jensen 3 , John McGuire 4, Yi Zheng 1, Carla Coltharp 1, Mei Jiang 5, Justin Lucas 6, Edwin Parra 5, Ignacio Wistuba 5, Darren Locke 6, Bernard A. Fox 3, David L. Rimm 4, Janis M. Taube 2
1 Akoya Biosciences, Hopkinton, MA, USA
2 The Johns Hopkins Hospital, Baltimore, MD, USA
3 Earle A. Chiles Research Institute, Portland, OR, USA
4 Yale University School of Medicine, New Haven, CT, USA
5 The University of Texas MD Anderson Cancer Center, Houston, TX, USA
6 Bristol Myers Squibb, Princeton, NJ, USA
Issue: 2019 American Association of Cancer Research Trade Show
Introduction
Emerging data suggests that predictive biomarkers based on the spatial arrangement of multiple cell types in FFPE tissue sections will be an important component of precision medicine in immuneoncology. (1) Multiplexed immunofluorescence (mIF) facilitates such assessments. If mIF is to play a translational role in research and ultimately clinical practice, it is vital to refine, standardize, and validate an end-to-end workflow that supports large scale multi-site trials and clinical laboratory processes. To this end, six institutions collaborated to develop an automated 6-plex assay focused on the PD-1/PD-L1 axis and assessed its inter- and intra-site reproducibility. Specific attention was paid to assessment of %PD-L1 expression by immune cells (ICs), as pathologists have poor concordance for this parameter. (2,3)
Methods
A 7-color mIF panel (PD-L1, PD-1, CD8, CD68, FoxP3, Cytokeratin, and DAPI) was optimized on a Leica Bond Rx autostainer. Serial sections of tonsil and a lung cancer tissue-microarray (TMA), antibodies and TSA-Opal detection reagents (Akoya Biosciences) were distributed to each site. Cell pellet arrays were also distributed and used to normalize batch variation in intensity measurements. Tonsil and TMA sections were stained at each site and imaged at 20x using a Vectra Polaris. Cells were segmented and phenotyped using image analysis algorithms. In tonsil sections, the average intensity of the top quartile of cells positive for each marker was assessed to identify potential variation in staining intensity. In lung TMAs, cell densities and %PD-L1 expression in immune cells (CD68+ and CD8+ cells) was determined.
Results
- The average staining intensity coefficients of variation (CV) for all markers within sites was 10% in tonsil samples.
- Inter-site concordance for tumor cell and immune cell subset densities in TMAs had an average R2 value of 0.86 and slope of 0.96.
- Inter-site concordance for %PD-L1+ ICs had an average R2 value of 0.81, in contrast to inter-class concordance values of <0.3 in the NCCN2 and Blueprint 2 studies. (3)
1. Lu, et al. SITC 2. Rimm DL, Han G, Taube JM, Yi ES, Bridge JA, Flieder DB, et al. A Prospective, Multi-institutional, Pathologist-Based Assessment of 4 Immunohistochemistry Assays for PD-L1 Expression in Non-Small Cell Lung Cancer. JAMA Oncol. 2017;3(8):1051-8. 3. Hirsch et. al., PD-L1 Immunohistochemistry Assays for Lung Cancer: Results from Phase 1 of the Blueprint PD-L1 IHC Assay Comparison Project, Feb. 2017, Journal of Thoracic Oncology, Vol. 12 No. 2: 208-222
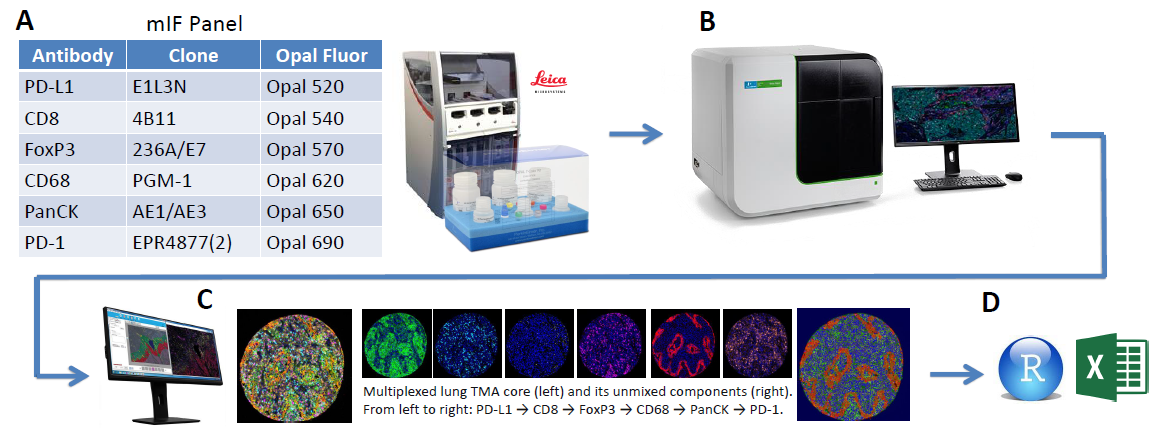
Figure 1. End-to-end workflow
(A) Staining was performed on the Leica Bond RX using the above mIF panel
(B) Multispectral slide imaging was performed on the Vectra Polaris (Akoya Biosciences, Hopkinton, MA)
(C) Image analysis with inForm software
(D) Data analysis using R and Excel
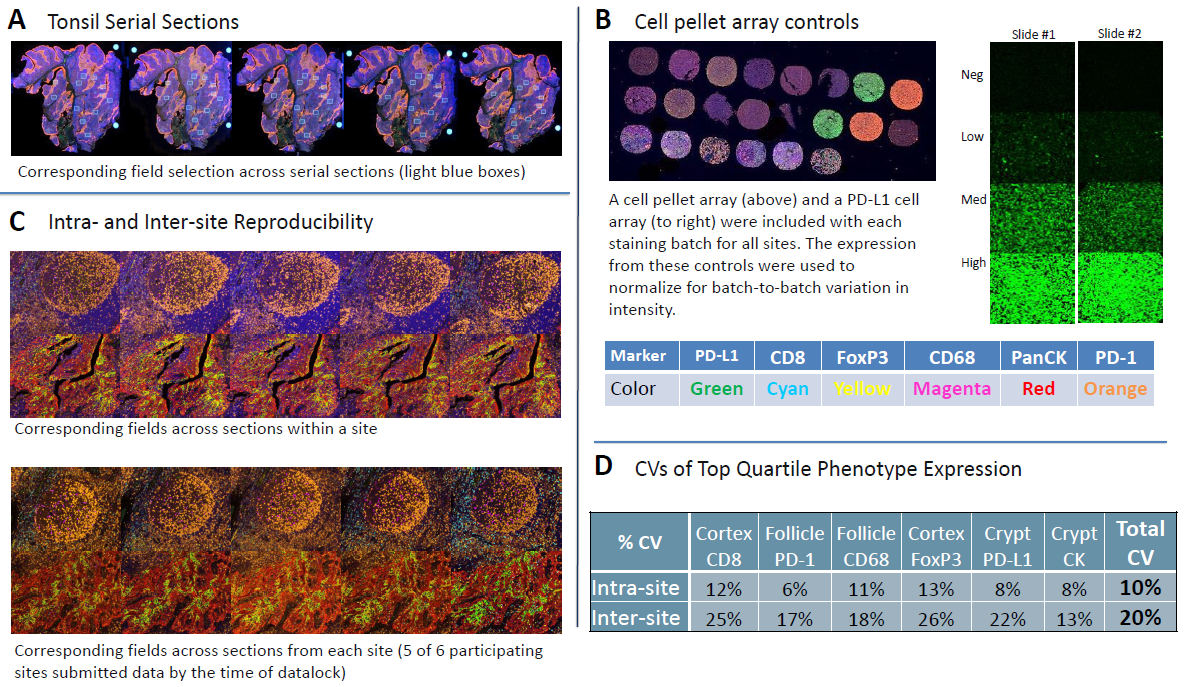
Figure 2. Reproducibility assessment with tonsil serial sections.
(A) Field selection across serial sections. 12 20x fields were selected per sample to have fields enriched for markers of interest (4 each from follicle, mantel, and cortex).
(B) Cell pellet arrays were used to normalize for batch-to-batch differences.
(C) Representative images from intra-site and inter-site analysis, showing corresponding fields from within site serial sections and site-to-site, respectively.
(D) Intra- and inter-site CVs calculated from the top quartile of expression for a given marker in a specific anatomic location.
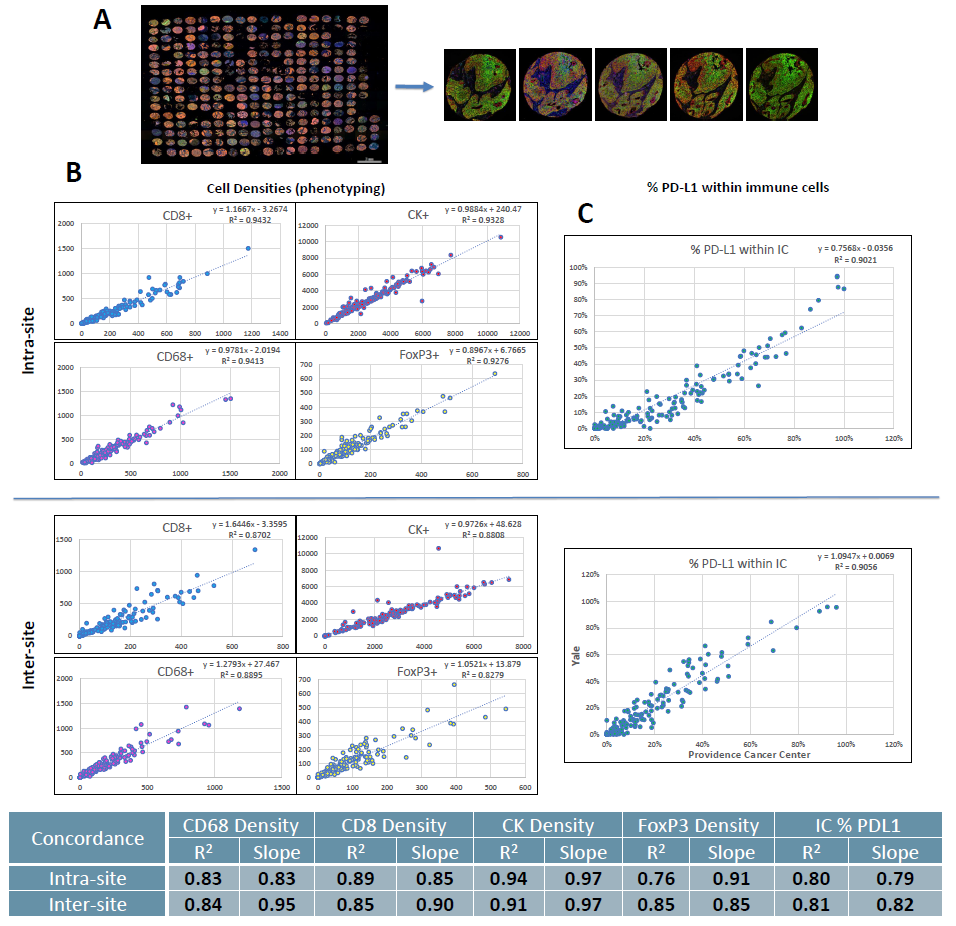
Figure 3. Correlation between adjacent lung TMA sections.
(A) Whole slide scan of one lung TMA and representative 20x fields from 1 core across sites.
(B) Example concordance plots of phenotype densities for each marker.
(C) Intra- and inter-site average concordance R2 and slope values.
Conclusions
- We demonstrate a reproducible end-to-end process for mIF characterization of the PD-1/PD-L1 axis including automated staining, multispectral imaging, and machine-learning-trained image analysis
- This approach improved reproducibility of %PD-L1 IC assessment and brought it in line with %PD-L1 tumor cell assessment by pathologists
- The described approach may serve as a template for assessing reproducibility of emerging mIF panels for other investigative teams, with an eye toward translating such approaches into clinical trials and ultimately into the clinic
